Rfam Uses
Rfam Uses - Search results - Wiki Rfam Uses
The page "Rfam+Uses" does not exist. You can create a draft and submit it for review or request that a redirect be created, but consider checking the search results below to see whether the topic is already covered.
- Rfam is a database containing information about non-coding RNA (ncRNA) families and other structured RNA elements. It is an annotated, open access database...
- G-test (section Distribution and use)computational linguistics community where it is now widely used. The R-scape program (used by Rfam) uses G-test to detect co-variation between RNA sequence alignment...
- (July 2015). "RNAiFold 2.0: a web server and software to design custom and Rfam-based RNA molecules". Nucleic Acids Research. 43 (W1): W513–W521. arXiv:1505...
 element 1 at Rfam Page for SECIS element 2 at Rfam Page for SECIS element 3 at Rfam Page for SECIS element 4 at Rfam Page for SECIS element 5 at Rfam...
element 1 at Rfam Page for SECIS element 2 at Rfam Page for SECIS element 3 at Rfam Page for SECIS element 4 at Rfam Page for SECIS element 5 at Rfam...- Stockholm format is a multiple sequence alignment format used by Pfam, Rfam and Dfam, to disseminate protein, RNA and DNA sequence alignments. The alignment...
 Ribonuclease P (category Rfam pages needing a picture)Nuclear RNase P at Rfam Page for Archaeal RNase P at Rfam Page for Bacterial RNase P class A at Rfam Page for Bacterial RNase P class B at Rfam RNase+P at the...
Ribonuclease P (category Rfam pages needing a picture)Nuclear RNase P at Rfam Page for Archaeal RNase P at Rfam Page for Bacterial RNase P class A at Rfam Page for Bacterial RNase P class B at Rfam RNase+P at the...- 18S ribosomal RNA (section Uses in phylogeny)easy to access due to highly conserved flanking regions allowing for the use of universal primers. Their repetitive arrangement within the genome provides...
- RNA Biology (category Use dmy dates from August 2022)a collaboration between the journal and the consortium that produces the Rfam database of RNA families. The journal is abstracted and indexed in: MEDLINE/PubMed...
 CRISPR (section Use by phages)Secondary structure taken from the Rfam database. Family RF01315. CRISPR-DR5: Secondary structure taken from the Rfam database. Family RF011318. CRISPR-DR6:...
CRISPR (section Use by phages)Secondary structure taken from the Rfam database. Family RF01315. CRISPR-DR5: Secondary structure taken from the Rfam database. Family RF011318. CRISPR-DR6:... required for DNA packaging". J. Biol. Chem. 265 (36): 22365–22370. doi:10.1016/S0021-9258(18)45714-6. PMID 2125049. Page for Bacteriophage pRNA at Rfam v t e...
required for DNA packaging". J. Biol. Chem. 265 (36): 22365–22370. doi:10.1016/S0021-9258(18)45714-6. PMID 2125049. Page for Bacteriophage pRNA at Rfam v t e... using a new RNA primer for each Okazaki fragment it encounters. Overall, the leading strand only uses one RNA primer, while the lagging strand uses a...
using a new RNA primer for each Okazaki fragment it encounters. Overall, the leading strand only uses one RNA primer, while the lagging strand uses a... Pfam (category Use dmy dates from April 2017)org), using duplicate independent data centres. This allowed for better centralisation of updates, and grouping with other Xfam projects such as Rfam, TreeFam...
Pfam (category Use dmy dates from April 2017)org), using duplicate independent data centres. This allowed for better centralisation of updates, and grouping with other Xfam projects such as Rfam, TreeFam... EntrezGene page for TERC Page for Vertebrate telomerase RNA at Rfam Page for Ciliate telomerase RNA at Rfam Page for Saccharomyces telomerase at Rfam...
EntrezGene page for TERC Page for Vertebrate telomerase RNA at Rfam Page for Ciliate telomerase RNA at Rfam Page for Saccharomyces telomerase at Rfam... "The uRNA database". Nucleic Acids Research. 25 (1): 102–3. doi:10.1093/nar/25.1.102. PMC 146409. PMID 9016512. Page for U6 spliceosomal RNA at Rfam...
"The uRNA database". Nucleic Acids Research. 25 (1): 102–3. doi:10.1093/nar/25.1.102. PMC 146409. PMID 9016512. Page for U6 spliceosomal RNA at Rfam... in Caenorhabditis elegans". BMC Molecular Biology. 8: 86. doi:10.1186/1471-2199-8-86. PMC 2200864. PMID 17903271. Page for SmY spliceosomal RNA at Rfam...
in Caenorhabditis elegans". BMC Molecular Biology. 8: 86. doi:10.1186/1471-2199-8-86. PMC 2200864. PMID 17903271. Page for SmY spliceosomal RNA at Rfam... Signal recognition particle RNA (category Rfam pages needing a picture)3-D models Rfam entry for Metazoan type signal recognition particle RNA Rfam entry for Bacterial small signal recognition particle RNA Rfam entry for Bacterial...
Signal recognition particle RNA (category Rfam pages needing a picture)3-D models Rfam entry for Metazoan type signal recognition particle RNA Rfam entry for Bacterial small signal recognition particle RNA Rfam entry for Bacterial...- Cis-regulatory element (category Use dmy dates from February 2021)Cis-trans isomerism Gene regulatory network Operon Promoter Trans-acting factor Rfam Transterm Davidson EH (2006). The Regulatory Genome: Gene Regulatory Networks...
- Probabilistic context-free grammar (section Example: Using evolutionary information to guide structure prediction)The RNA analysis package Infernal uses such profiles in inference of RNA alignments. The Rfam database also uses CMs in classifying RNAs into families...
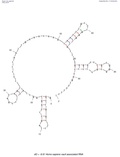 PMID 23871666. Stadler, Peter F. RNA Gene Prediction (PDF). Page for Vault RNA at Rfam The Vault website Vault+Ribonucleoprotein+Particles at the U.S. National...
PMID 23871666. Stadler, Peter F. RNA Gene Prediction (PDF). Page for Vault RNA at Rfam The Vault website Vault+Ribonucleoprotein+Particles at the U.S. National...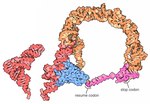 model built with both jakobid and oomycete sequences is now available at Rfam under the name ‘mt-tmRNA’. The standard bacterial tmRNA consists of a tRNA(Ala)-like...
model built with both jakobid and oomycete sequences is now available at Rfam under the name ‘mt-tmRNA’. The standard bacterial tmRNA consists of a tRNA(Ala)-like...











