Rfam Methods
Rfam Methods - Search results - Wiki Rfam Methods
The page "Rfam+Methods" does not exist. You can create a draft and submit it for review or request that a redirect be created, but consider checking the search results below to see whether the topic is already covered.
- Rfam is a database containing information about non-coding RNA (ncRNA) families and other structured RNA elements. It is an annotated, open access database...
- Stockholm format is a multiple sequence alignment format used by Pfam, Rfam and Dfam, to disseminate protein, RNA and DNA sequence alignments. The alignment...
 Ribonuclease P (category Rfam pages needing a picture)Nuclear RNase P at Rfam Page for Archaeal RNase P at Rfam Page for Bacterial RNase P class A at Rfam Page for Bacterial RNase P class B at Rfam RNase+P at the...
Ribonuclease P (category Rfam pages needing a picture)Nuclear RNase P at Rfam Page for Archaeal RNase P at Rfam Page for Bacterial RNase P class A at Rfam Page for Bacterial RNase P class B at Rfam RNase+P at the...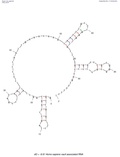 Vault RNA (section Research methods)PMID 23871666. Stadler, Peter F. RNA Gene Prediction (PDF). Page for Vault RNA at Rfam The Vault website Vault+Ribonucleoprotein+Particles at the U.S. National...
Vault RNA (section Research methods)PMID 23871666. Stadler, Peter F. RNA Gene Prediction (PDF). Page for Vault RNA at Rfam The Vault website Vault+Ribonucleoprotein+Particles at the U.S. National... Secondary structure taken from the Rfam database. Family RF01315. CRISPR-DR5: Secondary structure taken from the Rfam database. Family RF011318. CRISPR-DR6:...
Secondary structure taken from the Rfam database. Family RF01315. CRISPR-DR5: Secondary structure taken from the Rfam database. Family RF011318. CRISPR-DR6:...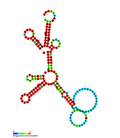 Methods in Molecular Biology. 999: 223–30. doi:10.1007/978-1-62703-357-2_16. PMID 23666702. Page for Human herpesvirus 1 small nucleolar RNA at Rfam Page...
Methods in Molecular Biology. 999: 223–30. doi:10.1007/978-1-62703-357-2_16. PMID 23666702. Page for Human herpesvirus 1 small nucleolar RNA at Rfam Page... TRAP regulatory protein". J Biol Chem. 277 (12): 10608–10613. doi:10.1074/jbc.M111813200. PMID 11786553. Page for Tryptophan operon leader at Rfam v t e...
TRAP regulatory protein". J Biol Chem. 277 (12): 10608–10613. doi:10.1074/jbc.M111813200. PMID 11786553. Page for Tryptophan operon leader at Rfam v t e... EntrezGene page for TERC Page for Vertebrate telomerase RNA at Rfam Page for Ciliate telomerase RNA at Rfam Page for Saccharomyces telomerase at Rfam...
EntrezGene page for TERC Page for Vertebrate telomerase RNA at Rfam Page for Ciliate telomerase RNA at Rfam Page for Saccharomyces telomerase at Rfam...- linguistics community where it is now widely used. The R-scape program (used by Rfam) uses G-test to detect co-variation between RNA sequence alignment positions...
- Cis-trans isomerism Gene regulatory network Operon Promoter Trans-acting factor Rfam Transterm Davidson EH (2006). The Regulatory Genome: Gene Regulatory Networks...
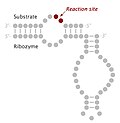 doi:10.1042/BST0301105. PMID 12440983. S2CID 32576742. Page for Hairpin ribozyme at Rfam Page for Hairpin-meta1 at Rfam Page for Hairpin-meta2 at Rfam...
doi:10.1042/BST0301105. PMID 12440983. S2CID 32576742. Page for Hairpin ribozyme at Rfam Page for Hairpin-meta1 at Rfam Page for Hairpin-meta2 at Rfam...- Mir-154 microRNA precursor family (category Rfam pages needing a picture)Developmental Dynamics. 236 (2): 572–80. doi:10.1002/dvdy.21047. PMC 2582151. PMID 17191223. Page for mir-154 microRNA precursor family at Rfam v t e...
- package Infernal uses such profiles in inference of RNA alignments. The Rfam database also uses CMs in classifying RNAs into families based on their structure...
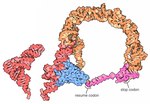 model built with both jakobid and oomycete sequences is now available at Rfam under the name ‘mt-tmRNA’. The standard bacterial tmRNA consists of a tRNA(Ala)-like...
model built with both jakobid and oomycete sequences is now available at Rfam under the name ‘mt-tmRNA’. The standard bacterial tmRNA consists of a tRNA(Ala)-like...- lab [1] Eric Westhof lab [2] Archived 2009-07-24 at the Wayback Machine Steinar Johansen lab [3] RNA catalysis Page for GIR1 branching ribozyme at Rfam...
 programma è il segmento di DNA compreso tra il primer forward e il primer reverse, retrieved 2021-11-30 Page for HIV primer binding site (PBS) at Rfam v t e...
programma è il segmento di DNA compreso tra il primer forward e il primer reverse, retrieved 2021-11-30 Page for HIV primer binding site (PBS) at Rfam v t e... T-box leader (section Method of regulation)will then form, thereby terminating transcription. Page for T-box leader at Rfam TBDB, a database of annotated T-box leader sequences Grigg JC, Chen Y, Grundy...
T-box leader (section Method of regulation)will then form, thereby terminating transcription. Page for T-box leader at Rfam TBDB, a database of annotated T-box leader sequences Grigg JC, Chen Y, Grundy... proposed from phylogenetic methods applied years ago between 2012 and 2013. Since then, a variety of novel phylogenetic methods have been proposed for Archaea...
proposed from phylogenetic methods applied years ago between 2012 and 2013. Since then, a variety of novel phylogenetic methods have been proposed for Archaea...- selection of pairwise alignments of between 30% and 40% sequence identity from Rfam (database) version 5.0. These parameters however, are not always effective...
 Journal of Molecular Biology. 310 (1): 111–126. doi:10.1006/jmbi.2001.4745. PMID 11419940. Page for c-myc internal ribosome entry site (IRES) at Rfam v t e...
Journal of Molecular Biology. 310 (1): 111–126. doi:10.1006/jmbi.2001.4745. PMID 11419940. Page for c-myc internal ribosome entry site (IRES) at Rfam v t e...











